Exploiting genomic data to understand viral evolution

An unusual algal virus
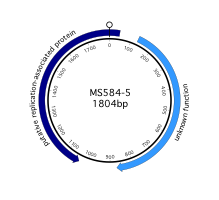
We isolated a high copy number viral genome from an algal cell whose genome was sequenced with the phi29 polymerase (Yoon et al., 2011). Some of its genomic architecture resembles a geminivirus (an ambisense geneome with a putative Rep gene in the complementary sense), but also has strong similarities to characterized animal circoviruses: a similar stem-loop including an identical origin of replication (CAGTATTAC) to canary circovirus and a basic residue-rich N-terminus to the predicted protein product of the unknown ORF, just like circovirus coat proteins. However, the putative replication-associated protein has the greatest similarity to Rep proteins of nanoviruses, a third family of ssDNA viruses. The novel genome has no homology with known algal viruses, and currently defies classification.
Novel ssDNA virus discovery
The recent commercialization of a polymerase (isolated from phage phi29) that preferentially amplifies circular single-stranded DNA has led to a new age in the discovery and taxonomy of ssDNA viruses, most of which have small, circular genomes. We and our collaborators are searching for novel ssDNA viruses both where we expect to find them them (in the insect vectors that transmit plant pathogenic ssDNA geminiviruses) and in metagenomic data sets. We are very interested in understanding how various ssDNA virus families are related to one another and assessing the ease of gene flow between these distinct kinds of ssDNA viruses.
RNA viruses evolve quickly and RNA viruses are small (always less than ~32 kb, usually less than ~15 kb). Both of these traits are attributed to the high mutation rate of RNA viruses, which itself is attributed to the lack of proofreading of RNA polymerases and the inability of error-correction enzymes to operate on RNA. However, there are other kinds of size-limited viruses, such as ssDNA viruses; none larger than ~25kb have been found. Through our research and the work of other groups, we now know many ssDNA viruses evolve as quickly as RNA viruses. How do ssDNA viruses have similar sizes and similar evolutionary dynamics as RNA viruses when they are not replicated with RNA polymerases. Do ssDNA viruses have high mutation rates, and if so, how do they achieve them when they are replicated with host cellular DNA polymerases? Are ssDNA viruses size limited for the similar reasons as the similarly sized RNA viruses? We use both experimental and bioinformatic approaches to this question.
We use a Bayesian MCMC phylogenetics program called BEAST to analyze viral sequences sampled from nature at different times to estimate the rate of molecular evolution in RNA and ssDNA viruses. It calculates a distribution of the most likely nucleotide substitution rate: the number of fixed mutations (substitutions) per site per year. If the evolution is completely neutral, then the substitution rate only reflects the mutation rate, so organisms that have a high neutral substitution rate must have a high mutation rate. If there is substantial positive selection -- when a virus is adapting to a novel host, perhaps -- then the substitution rate can be higher than the neutral substitution rate. Conversely, if there is purifying selection, as occurs in important protein-coding genes, the the substitution rate will be lower than the neutral substitution rate as most mutations are selected against and lost from the population. While viral sequences in GenBank don’t immediately tell us anything about the mutation rate of their species, nucleotide substitution rates offers us a window onto mutation rates and rooted phylogenies contain some information on mutational dynamics. These results inform the design of our wet lab experiments on fast-evolving viral population diversity and mutation rates.